
Concept explainers
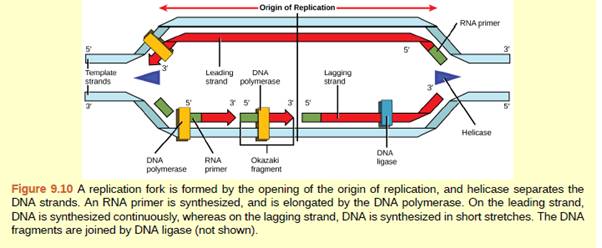
Figure 9.10 You isolate a cell strain in which the joining together of Okazaki fragments is impaired and suspect that a mutation has occurred in an enzyme found at the replication fork. Which enzyme is most likely to be mutated?
To write:
The enzyme which is most likely mutated when joining of Okazaki fragments is impaired in the replication.
Introduction:
DNA replication is the process in which the copy of DNA is made. This is done in 3 steps in a eukaryotic cell. They are initiation, elongation, and termination. It is a semiconservative type of replication. Enzymes involved in this process are helicase, DNA polymerase, and DNA ligase.
Explanation of Solution
DNA replication is started at the origin of replication where a Y shaped fork known as replication is formed when both strands are separated by helicase enzyme. Both the strands of DNA act as a template. One strand is known as a leading strand at which replication is continuous and other strand is known as a lagging strand at which replication occurs in fragments. This is because DNA polymerase can synthesize DNA only in 5' to 3' direction. These fragments are known as Okazaki fragments. These fragments at the lagging strands are then sealed by the enzyme DNA ligase. The joining of Okazaki fragments is impaired when the DNA ligase is not able to join the strands. This can occur due to the mutation in enzyme DNA ligase.
When joining of DNA fragments on the lagging strand is impaired in DNA replication, the mutation is most likely to occur in DNA ligase as it is an important enzyme which joins the strands.
Want to see more full solutions like this?
Chapter 9 Solutions
Concepts of Biology
Additional Science Textbook Solutions
College Physics
Human Biology: Concepts and Current Issues
Anatomy & Physiology
Essentials of Genetics (9th Edition) - Standalone book
Campbell Biology in Focus
Microbiology with Diseases by Body System (4th Edition)
- Human Fbh1 helicase is important in the process of DNA replication. When a mutation occurs during the production of Fbh1, the result is a mutant Fbh1 that binds at the replication fork and prevents any helicase protein from attaching to the strand. Based on this information and the image shown, what would happen during DNA replication if this mutant helicase were present? A - Topoisomerase would unwind the DNA and an RNA primer would attach to the DNA molecule and initiate replication. The process would then stop at the blue triangle because helicase is needed to separate the strands of DNA. B - Topoisomerase would unwind the DNA, but then the process would stop at the blue triangle because helicase, the RNA primer, would not be able to attach to the DNA molecule and initiate replication. C - The process would begin at the blue triangle when topoisomerase unwinds the DNA and an RNA primer attaches to the DNA molecule and initiates replication. DNA polymerase would begin the synthesis…arrow_forwardShown below is a long template strand of DNA where lagging strand DNA synthesis is occurring. The short horizontal lines represent two Okazaki fragments that have already been made. In the context of the replication fork, select the letter(d–g) that indicates where primase will synthesize the next RNA primer. Explain why did you choose that location?arrow_forwardA conditional mutation expresses its mutant phenotype only under certain conditions (the restrictive conditions) and expresses the normal phenotype under other conditions (the permissive conditions). One type of conditional mutation is a temperature-sensitive mutation, which expresses the mutant phenotype only at certain temperatures. Strains of E. coli have been isolated that contain temperature-sensitive mutations in the genes encoding different components of the replication machinery. In each of these strains, the protein produced by the mutated gene is non-functional under the restrictive conditions. These strains are grown under permissive conditions and then abruptly switched to the restrictive condition. After one round of replication under the restrictive condition, the DNA from each strain is isolated and analysed. What characteristics would you expect to see in the DNA isolated from each strain with a temperature-sensitive mutation in its gene that encodes a) DNA…arrow_forward
- Shown below is a drawing showing the result of an experiment in which an RNA molecule is allowed to mix with genomic DNA that has been denatured by boiling, and the two molecules are allowed to hybridize. The DNA strand is presumed to be the lighter-shaded one on the top. Note that only one strand of DNA is shown. This result was the first evidence for which of the following processes? a Replication b Transcription c Translation d Splicingarrow_forwardA genetics instructor designs a laboratory experiment to study the effects of UV radiation on mutation in bacteria. In the experiment, the students spread bacteria on petri plates, expose the plates to UV light for different lengths of time, place the plates in an incubator for 48 hours, and then count the number of colonies that appear on each plate. The bacteria that have received more UV radiation should have more pyrimidine dimers, which block replication; thus, fewer colonies should appear on the plates exposed to UV light for longer periods. Before the students carry out the experiment, the instructor warns them that while the bacteria are in the incubator, the students must not open the incubator door unless the room is darkened. Why should the bacteria not be exposed to light?arrow_forwardA conditional mutation expresses its mutant phenotype only under certain conditions (the restrictive conditions) and expresses the normal phenotype under other conditions (the permissive conditions). One type of conditional mutation is a temperature-sensitive mutation, which expresses the mutant phenotype only at certain temperatures. Strains of E. coli have been isolated that contain temperature-sensitive mutations in genes encoding different components of the replication machinery. In each of these strains, the protein produced by the mutated gene is nonfunctional under the restrictive conditions. You grow these strains under the permissive conditions and then abruptly switch them to the restrictive conditions. After one round of replication under therestrictive conditions, you isolate DNA from each strain and analyze it. What characteristics would you expect to see in the DNA isolated from a strain with a temperature-sensitive mutation in the gene that encodes the following…arrow_forward
- A conditional mutation expresses its mutant phenotype only under certain conditions (the restrictive conditions) and expresses the normal phenotype under other conditions (the permissive conditions). One type of conditional mutation is a temperature-sensitive mutation, which expresses the mutant phenotype only at certain temperatures. Strains of E. coli have been isolated that contain temperature-sensitive mutations in genes encoding different components of the replication machinery. In each of these strains, the protein produced by the mutated gene is nonfunctional under the restrictive conditions. You grow these strains under the permissive conditions and then abruptly switch them to the restrictive conditions. After one round of replication under therestrictive conditions, you isolate DNA from each strain and analyze it. What characteristics would you expect to see in the DNA isolated from a strain with a temperature-sensitive mutation in the gene that encodes the following…arrow_forwardA conditional mutation expresses its mutant phenotype only under certain conditions (the restrictive conditions) and expresses the normal phenotype under other conditions (the permissive conditions). One type of conditional mutation is a temperature-sensitive mutation, which expresses the mutant phenotype only at certain temperatures. Strains of E. coli have been isolated that contain temperature-sensitive mutations in genes encoding different components of the replication machinery. In each of these strains, the protein produced by the mutated gene is nonfunctional under the restrictive conditions. You grow these strains under the permissive conditions and then abruptly switch them to the restrictive conditions. After one round of replication under therestrictive conditions, you isolate DNA from each strain and analyze it. What characteristics would you expect to see in the DNA isolated from a strain with a temperature-sensitive mutation in the gene that encodes the following…arrow_forwardA conditional mutation expresses its mutant phenotype only under certain conditions (the restrictive conditions) and expresses the normal phenotype under other conditions (the permissive conditions). One type of conditional mutation is a temperature-sensitive mutation, which expresses the mutant phenotype only at certain temperatures. Strains of E. coli have been isolated that contain temperature-sensitive mutations in genes encoding different components of the replication machinery. In each of these strains, the protein produced by the mutated gene is nonfunctional under the restrictive conditions. You grow these strains under the permissive conditions and then abruptly switch them to the restrictive conditions. After one round of replication under the restrictive conditions, you isolate DNA from each strain and analyze it. What characteristics would you expect to see in the DNA isolated from a strain with a temperature-sensitive mutation in the gene that encodes each of the following…arrow_forward
- You conducted an experiment to determine the mechanism of DNA replication in the hypothetical organism Fungus mungus. Your data shows that synthesis of newly replicated DNA from F. mungus is discontinuous on both strands of the replication fork. Does this result support or not support the hypothesis that F. mungus replicates its DNA by the same mechanism as yeast? Briefly explain your answer.arrow_forwardIn the diagram below, the arrow indicates the direction of movement of a replication fork. Replication fork Unwinding (a) Which template strand (top or bottom) is the lead strand? (b) Which strand (top or bottom) would you expect to find the Okazaki fragments? Bottom; Top Bottom; Bottom O Top; Top O Top; Bottomarrow_forwardWhether the statement "In E. coli, where the replication fork travels at 500 nucleotide pairs per second, the DNA ahead of the fork-in the absence of topoisomerase-would have to rotate at nearly 3000 revolutions per minute" is true or false.arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





